Taxonomy and Phylogeny
Taxonomy of the bacteria was historically based on phenotypical physical and chemical characteristics – phenetic taxonomy. According to "Bergey's Manual of Systematic Bacteriology", all bacteria can be classified into four divisions, or phyla according to the constituents of their cell walls. [flow chart for classification of obligate intracellular bacteria]
Each division was further subdivided into sections according to -
 1. Reaction to the Gram stain due to thick layer of peptidoglycan (Gram +ve) or thin layer of peptidoglycan (Gram -ve). The cell walls of Archaeobacteria contain no peptidoglycan.
1. Reaction to the Gram stain due to thick layer of peptidoglycan (Gram +ve) or thin layer of peptidoglycan (Gram -ve). The cell walls of Archaeobacteria contain no peptidoglycan.
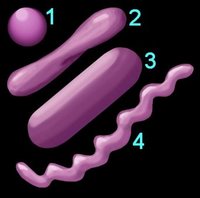
2. Shape (left) – cocci (1), pleomorphic (2), bacillus (3), helical (4) 3. Arrangement of cells
4. Oxygen requirement – aerobic, anaerobic, or facultative anaerobe
5. Motility – flagellated, non-motile
6. Specific nutritional requirements
7. Trophic (metabolic) properties – autotrophic (chemical or photosynthetic), heterotrophic
The major subdivisions employed in general taxonomies are:
Domain, Kingdom, Phylum, Class, Order, Family, Genus, and Species. The mnemonic "Dashing King Philip Came Over From Greater Spain" applies.
These subdivisions may be further subdivided. It is common for bacteria to be subdivided into Divisions and further subdivided into Orders. The advent of genomics and examination of 16s ribosomal RNA has led to modifications in the classification of the prokaryotes, with the older nomenclatures revised into new classifications of Phyla.
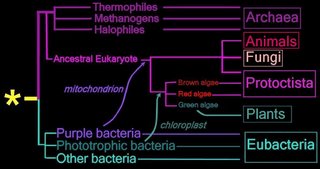 Right - click to enlarge image: Proposed endosymbiotic transfer events between the three Domains and the six Kingdoms of Life. (below) Both the Eubacteria and Archaea are prokaryotes, while animals, fungi, plants, and protists are eukaryotes.
Right - click to enlarge image: Proposed endosymbiotic transfer events between the three Domains and the six Kingdoms of Life. (below) Both the Eubacteria and Archaea are prokaryotes, while animals, fungi, plants, and protists are eukaryotes.
The yellow asterisk * indicates the last universal common ancestor (LUCA), or universal cenancestor, which is hypothesized as being at the ancestral root of all living organisms. Not the earliest or simplest living organism, and not necessarily the sole example of its type, this organism possessed the genetic material that diverged (about 3.5 Ga) into all current living organisms. A number of terms are employed to refer to the universal cenancestor – last universal ancestor (LUA), last common ancestor (LCA), or last universal common ancestor (LUCA).
Woese and Fox proposed the Three Domain system: Eubacteria, Archaea, and Eukaryotes.
The Five Kingdom system was proposed in 1969: Monera (prokaryotes), Protista, Plantae, Fungi, Animalia. Discovery of the Archaea added the sixth kingdom.
History of taxonomic concepts:
Linnaeus, 1735 – 2 Kingdoms – Animalia, Vegetabilia
Haeckel, 1866 – 3 Kingdoms – Protista, Plantae, Animalia. Image Haeckel's tree of life.
Chatton, 1937 – 2 Empires – Prokaryota, Eukaryota
Copeland, 1956 – 4 Kingdoms – Monera (prokaryotes), Protoctista, Plantae, Animalia
Whittaker, 1969 – Monera (prokaryotes), Fungi, Protista, Plantae, Animalia
Woese et al, 1977 – 6 Kingdom – Eubacteria, Archaea, Protista, Fungi, Plantae, Animalia
Woese and Fox, 1999 – 3 Domain system: Eubacteria, Archaea, and Eukaryotes
When Woese and Fox proposed the 3 Domain system, the term 'Urkaryotes' was proposed for ancestors of eukaryotes prior to their endosymbiotic acquisition of mitochondria and chloroplasts from prokaryotes. Margulis and Schwartz proposed that Kingdom Protista be replaced by Protoctista to reflect inclusion of multicellular organisms that did fit into the other three eukaryotic kingdoms.
"Misunderstanding the Bacteriological Code"
The Three Domain system is increasingly accepted. Free Full Text Article : Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Woese CR, Fox GE. Proc Natl Acad Sci U S A. 1977 Nov;74(11):5088-90.
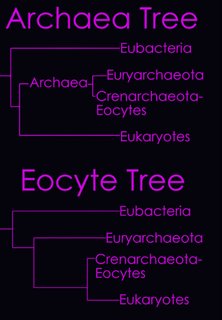
According to the Tree of Life Web Project, two alternative views on the relationship of the major lineages (omitting viruses) are currently regarded as viable (right - click to enlarge image).
 Left - click to enlarge image: Horizontal gene transfer - gene swapping - has blurred the evolutionary relationships (image) of prokaryotes, and continues to provide a mechanism for the sharing of antibiotic resistance between bacteria. See: Trees, vines and nets: Microbial evolution changes its face.
Left - click to enlarge image: Horizontal gene transfer - gene swapping - has blurred the evolutionary relationships (image) of prokaryotes, and continues to provide a mechanism for the sharing of antibiotic resistance between bacteria. See: Trees, vines and nets: Microbial evolution changes its face.
Phylogenetic separation into evolutionary relationships (clades), based on comparison of genomes is likely to supplant phenotypical (phenetic) taxonomies of the prokaryotes.
(nomenclature)
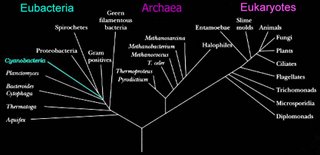 Left - click to enlarge image: Woese has proposed a scheme based on the 16s subunit of ribosomal RNA, which appears to better illustrate evolutionary relationships within the 3 domains.
Left - click to enlarge image: Woese has proposed a scheme based on the 16s subunit of ribosomal RNA, which appears to better illustrate evolutionary relationships within the 3 domains.
Phylogenetic tree of organisms
Domain combinations in archaeal, eubacterial and eukaryotic proteomes.
There is a limited repertoire of domain families that are duplicated and combined in different ways to form the set of proteins in a genome. Proteins are gene products, and at the level of genes, duplication, recombination, fusion and fission are the processes that produce new genes. We attempt to gain an overview of these processes by studying the evolutionary units in proteins, domains, in the protein sequences of 40 genomes. The domain and superfamily definitions in the Structural Classification of Proteins Database are used, so that we can view all pairs of adjacent domains in genome sequences in terms of their superfamily combinations. We find 783 out of the 859 superfamilies in SCOP in these genomes, and the 783 families occur in 1307 pairwise combinations. Most families are observed in combination with one or two other families, while a few families are very versatile in their combinatorial behaviour; 209 families do not make combinations with other families. This type of pattern can be described as a scale-free network. We also study the N to C-terminal orientation of domain pairs and domain repeats. The phylogenetic distribution of domain combinations is surveyed, to establish the extent of common and kingdom-specific combinations. Of the kingdom-specific combinations, significantly more combinations consist of families present in all three kingdoms than of families present in one or two kingdoms. Hence, we are led to conclude that recombination between common families, as compared to the invention of new families and recombination among these, has also been a major contribution to the evolution of kingdom-specific and species-specific functions in organisms in all three kingdoms. Finally, we compare the set of the domain combinations in the genomes to those in the RCSB Protein Data Bank, and discuss the implications for structural genomics.
Apic G, Gough J, Teichmann SA. Domain combinations in archaeal, eubacterial and eukaryotic proteomes. J Mol Biol. 2001 Jul 6;310(2):311-25.
Table Comparisons of Eubacteria, Archaea, and Eukaryotes Electron acceptors for respiration and methanogenesis in prokaryotes Glycolysis in bacteria Lithotrophic prokaryotes Comparison of plant and bacterial photosynthesis - The Three Domains View Quicktime Movie - Genetic Data movie of phylogram construction - image cladogram - image Tree of Life— Lateral Gene Transfer Diagram - image uprooted tree - image 16S ribosomal RNA - image The "Shrub of Life" - image A comparison of key characteristics from the three domains of life - enlarged - Genomics Animations and Images - Proteins & Proteomics - Animations and Images – Evolution and Phylogenetics - Animations and Images - Biodiversity - Animations and Images – Microbial Diversity – Animations and Images – Emerging Infectious Diseases - Animations and Images – HIV & AIDS - Animations and Images :
Each division was further subdivided into sections according to -
 1. Reaction to the Gram stain due to thick layer of peptidoglycan (Gram +ve) or thin layer of peptidoglycan (Gram -ve). The cell walls of Archaeobacteria contain no peptidoglycan.
1. Reaction to the Gram stain due to thick layer of peptidoglycan (Gram +ve) or thin layer of peptidoglycan (Gram -ve). The cell walls of Archaeobacteria contain no peptidoglycan.2. Shape (left) – cocci (1), pleomorphic (2), bacillus (3), helical (4) 3. Arrangement of cells
4. Oxygen requirement – aerobic, anaerobic, or facultative anaerobe
5. Motility – flagellated, non-motile
6. Specific nutritional requirements
7. Trophic (metabolic) properties – autotrophic (chemical or photosynthetic), heterotrophic
The major subdivisions employed in general taxonomies are:
Domain, Kingdom, Phylum, Class, Order, Family, Genus, and Species. The mnemonic "Dashing King Philip Came Over From Greater Spain" applies.
These subdivisions may be further subdivided. It is common for bacteria to be subdivided into Divisions and further subdivided into Orders. The advent of genomics and examination of 16s ribosomal RNA has led to modifications in the classification of the prokaryotes, with the older nomenclatures revised into new classifications of Phyla.
 Right - click to enlarge image: Proposed endosymbiotic transfer events between the three Domains and the six Kingdoms of Life. (below) Both the Eubacteria and Archaea are prokaryotes, while animals, fungi, plants, and protists are eukaryotes.
Right - click to enlarge image: Proposed endosymbiotic transfer events between the three Domains and the six Kingdoms of Life. (below) Both the Eubacteria and Archaea are prokaryotes, while animals, fungi, plants, and protists are eukaryotes.The yellow asterisk * indicates the last universal common ancestor (LUCA), or universal cenancestor, which is hypothesized as being at the ancestral root of all living organisms. Not the earliest or simplest living organism, and not necessarily the sole example of its type, this organism possessed the genetic material that diverged (about 3.5 Ga) into all current living organisms. A number of terms are employed to refer to the universal cenancestor – last universal ancestor (LUA), last common ancestor (LCA), or last universal common ancestor (LUCA).
Woese and Fox proposed the Three Domain system: Eubacteria, Archaea, and Eukaryotes.
The Five Kingdom system was proposed in 1969: Monera (prokaryotes), Protista, Plantae, Fungi, Animalia. Discovery of the Archaea added the sixth kingdom.
History of taxonomic concepts:
Linnaeus, 1735 – 2 Kingdoms – Animalia, Vegetabilia
Haeckel, 1866 – 3 Kingdoms – Protista, Plantae, Animalia. Image Haeckel's tree of life.
Chatton, 1937 – 2 Empires – Prokaryota, Eukaryota
Copeland, 1956 – 4 Kingdoms – Monera (prokaryotes), Protoctista, Plantae, Animalia
Whittaker, 1969 – Monera (prokaryotes), Fungi, Protista, Plantae, Animalia
Woese et al, 1977 – 6 Kingdom – Eubacteria, Archaea, Protista, Fungi, Plantae, Animalia
Woese and Fox, 1999 – 3 Domain system: Eubacteria, Archaea, and Eukaryotes
When Woese and Fox proposed the 3 Domain system, the term 'Urkaryotes' was proposed for ancestors of eukaryotes prior to their endosymbiotic acquisition of mitochondria and chloroplasts from prokaryotes. Margulis and Schwartz proposed that Kingdom Protista be replaced by Protoctista to reflect inclusion of multicellular organisms that did fit into the other three eukaryotic kingdoms.
"Misunderstanding the Bacteriological Code"
The Three Domain system is increasingly accepted. Free Full Text Article : Phylogenetic structure of the prokaryotic domain: The primary kingdoms. Woese CR, Fox GE. Proc Natl Acad Sci U S A. 1977 Nov;74(11):5088-90.

According to the Tree of Life Web Project, two alternative views on the relationship of the major lineages (omitting viruses) are currently regarded as viable (right - click to enlarge image).
 Left - click to enlarge image: Horizontal gene transfer - gene swapping - has blurred the evolutionary relationships (image) of prokaryotes, and continues to provide a mechanism for the sharing of antibiotic resistance between bacteria. See: Trees, vines and nets: Microbial evolution changes its face.
Left - click to enlarge image: Horizontal gene transfer - gene swapping - has blurred the evolutionary relationships (image) of prokaryotes, and continues to provide a mechanism for the sharing of antibiotic resistance between bacteria. See: Trees, vines and nets: Microbial evolution changes its face.Phylogenetic separation into evolutionary relationships (clades), based on comparison of genomes is likely to supplant phenotypical (phenetic) taxonomies of the prokaryotes.
(nomenclature)
 Left - click to enlarge image: Woese has proposed a scheme based on the 16s subunit of ribosomal RNA, which appears to better illustrate evolutionary relationships within the 3 domains.
Left - click to enlarge image: Woese has proposed a scheme based on the 16s subunit of ribosomal RNA, which appears to better illustrate evolutionary relationships within the 3 domains.Phylogenetic tree of organisms
Domain combinations in archaeal, eubacterial and eukaryotic proteomes.
There is a limited repertoire of domain families that are duplicated and combined in different ways to form the set of proteins in a genome. Proteins are gene products, and at the level of genes, duplication, recombination, fusion and fission are the processes that produce new genes. We attempt to gain an overview of these processes by studying the evolutionary units in proteins, domains, in the protein sequences of 40 genomes. The domain and superfamily definitions in the Structural Classification of Proteins Database are used, so that we can view all pairs of adjacent domains in genome sequences in terms of their superfamily combinations. We find 783 out of the 859 superfamilies in SCOP in these genomes, and the 783 families occur in 1307 pairwise combinations. Most families are observed in combination with one or two other families, while a few families are very versatile in their combinatorial behaviour; 209 families do not make combinations with other families. This type of pattern can be described as a scale-free network. We also study the N to C-terminal orientation of domain pairs and domain repeats. The phylogenetic distribution of domain combinations is surveyed, to establish the extent of common and kingdom-specific combinations. Of the kingdom-specific combinations, significantly more combinations consist of families present in all three kingdoms than of families present in one or two kingdoms. Hence, we are led to conclude that recombination between common families, as compared to the invention of new families and recombination among these, has also been a major contribution to the evolution of kingdom-specific and species-specific functions in organisms in all three kingdoms. Finally, we compare the set of the domain combinations in the genomes to those in the RCSB Protein Data Bank, and discuss the implications for structural genomics.
Apic G, Gough J, Teichmann SA. Domain combinations in archaeal, eubacterial and eukaryotic proteomes. J Mol Biol. 2001 Jul 6;310(2):311-25.
Table Comparisons of Eubacteria, Archaea, and Eukaryotes Electron acceptors for respiration and methanogenesis in prokaryotes Glycolysis in bacteria Lithotrophic prokaryotes Comparison of plant and bacterial photosynthesis - The Three Domains View Quicktime Movie - Genetic Data movie of phylogram construction - image cladogram - image Tree of Life— Lateral Gene Transfer Diagram - image uprooted tree - image 16S ribosomal RNA - image The "Shrub of Life" - image A comparison of key characteristics from the three domains of life - enlarged - Genomics Animations and Images - Proteins & Proteomics - Animations and Images – Evolution and Phylogenetics - Animations and Images - Biodiversity - Animations and Images – Microbial Diversity – Animations and Images – Emerging Infectious Diseases - Animations and Images – HIV & AIDS - Animations and Images :
Labels: phenetic taxonomy








































2 Glossary:
Taxonomies aim to group organisms according to shared characteristics against the background of biological diversity.
Phenetic system: groupings of organisms based on mutual similarity of phenotypic (physical and chemical) characteristics. Phenetic groupings may or may not correlate with evolutionary relationships.
Numerical Taxonomy: a common approach to phenetic taxonomy, which employs a number of phenotypic characteristics to generate similarity coefficients that may be mapped in dendrograms. Groupings based on numerical taxonomy may or may not correlate with evolutionary relationships.
Phylogenetic system: groups organisms based on shared evolutionary heritage. DNA and RNA sequencing techniques are considered to give the most meaningful phylogenies.
Monophyletic taxon or clade: an accurate grouping of only (opp. polyphyletic) and all (opp. paraphyletic) descendents of a shared common ancestor. A monopyletic group is genetically homogeneous and reflects evolutionary relationships.
Paraphyletic taxon or clade: a monophyletic group that excludes one or more discrete groups descended from the most recent common ancestral species of the entire group. Other descendent species of the most recent common ancestor have been excluded from the paraphyletic taxon, usually because of morphologic distinctiveness.
Polyphyletic taxon: opposite to monophyletic taxon: A polyphyletic group is mistakenly or improperly erected on the basis of homoplasy.—characteristics that have arisen despite not sharing a common ancestor. Homoplasy arises because of convergent evolution, parallelism, evolutionary reversals, horizontal gene transfer, or gene duplications. Polyphyletic taxa are genetically heterogeneous because members do not share a common ancestor.
Archaea constitute a recently recognized phylogenetic domain. While eubacteria and archaeobacteria are similar in structure, they have a different metabolism and genotype. Archaea can be distinguished from bacteria in that their cell walls do not have murein—a peptidoglycan-containing muramic acid. Their cell membranes are unique in including isopranyl ether lipids. A defining physiological characteristic of archaea is their ability to live in extreme environments. Thus, these organisms are called “extremophiles” and unlike eubacteria and eukarya depend environmental conditions such as high salinity, extremes of temperature, unusual chemical substrates, or high pressure.
Post a Comment
<< Home